IMSB News
All stories that have been tagged with IMSB Projects
Master/Semestert Project: Metabolome:Proteome Cross-Talk in Basal Autophagy
For students of D-BIOL/D-HEST/D-BSSE
Master/Semester Project: Complexity of Mitochondrial Transporters in KRAS-driven Colorectal Cancer
For students in D-BIOL/D-HEST/D-BSSE
Master/Semester Project: Complexity of mitochondrial transporters and metabolic rewiring III
For students of D-BIOL/D-HEST/D-BSSE
Master/Semester Project: Complexity of mitochondrial transporters and metabolic rewiring II
For students of D-BIOL/D-HEST/D-BSSE
Master/Semester Project: Complexity of mitochondrial transporters and metabolic rewiring I
For students of D-BIOL/D-HEST/D-BSSE
Master/Semester Project: Interfacing microfluidics and high-resolution mass spectrometry for the study of complex metabolomes II
For students of D-BSSE/D-MATL/D-CHAB/D-BIOL
Master/Semester Project: Master’s Student Project: Interfacing microfluidics and high-resolution mass spectrometry for the study of complex metabolomes I
For students of D-BSSE/D-MATL/D-CHAB/D-BIOL
Master/semester project - Prediction and characterization of unknown membrane protein function
In this project, we will systematically characterize the function of unknown membrane proteins and their role in the cell's response to environmental conditions.
Absolute stoichiometry determination of dynamic protein complexes
Mass spectrometry is the widely used accurate quantification of the proteome composition. However very few attempts have been made to use these techniques for the determination of the absolute stoichiometry of protein complexes. The goal of this project is to use Single Reaction Monitoring (SRM)-MS, in combination with AQUA peptides, to define the absolute stoichiometry of dynamically assembling protein complexes that are relevant in the context of inflammation and autoimmune diseases.
Master thesis in Wollscheid Lab
Mass Spectrometric Analysis of Hepatitis B Virus X Proteoforms and Interactomes.
Hafen Lab: Semester project and/or Master thesis on the genetics of inter-individual variations in human sensory perception
The project lies at the intersection of genetics, psychophysics of smell, and data democracy. Students will have the opportunity to combine wet lab work, study design, and computational analysis. Students should be able to write simple scripts in R, Matlab, or Python or have the willingness to acquire and develop these skills.
Sauer Lab: Master thesis "Investigating protein-metabolite interactions in E. coli"
In this project, we aim at inferring functional protein-metabolite interactions in the bacterium Escherichia coli from dynamic metabolomics data.
Hafen lab: Master’s thesis and/or semester project on the role of cryptic genetic variation on circadian behaviors
Cryptic genetic variation (CGV) is the hidden genetic variation accumulated over time that do not contribute to the normal range of phenotypes, but may provide a source of adaptive variation to novel conditions. This project will examine if the CGV in different genetic backgrounds is influential in the evolution of circadian traits. (Keywords: Drosophila melanogaster, genetics, behavior, circadian rhythms)
Hafen Lab: Master thesis/semester project on CRISPR/Cas9-mediated SNP changes to elucidate cause of size variations
How genetic variation (e.g. SNP) leads to different traits/phenotypes in natural populations is a fundamental and challenging problem in both the life and medical sciences. This project will involve several objectives to tackle this problem by first establishing an efficient in vivo CRISPR/Cas9 method and then using this method to change phenotype-associated SNPs in inbred Drosophila lines derived from a natural population.
Sauer group: Interplay of posttranslational modification sites in enzyme activity regulation (Master project)
Research project available on investigating the interplay and protein activity fine-tuning of different posttranslational modifications (PTM) sites
Sauer group: Metabolomics and Flux Balance Analysis in Saccharomyces cerevisiae (Semester/Master Project)
Research project available on investigating the regulation of metabolism in S. cerevisiae, via enzyme phosphorylation
Zamboni group, Metabolomics & network analysis
Research project available on investigating metabolic response to mitochndrial defects in yeast
Aebersold group: Student project to monitor epigenetic states in melanoma cells using novel proteomic tools
The student will learn how to process samples for mass spectrometry analysis, to analyse the resulting proteomic datasets and integrate them into a larger multi-omics framework. The work will be published in a scientific journal and the student will be involved in writing up the manuscript.
Master thesis in Wollscheid Lab
Mass Spectrometric Analysis of Hepatitis B Virus X Proteoforms and Interactomes.
Stocker lab, Master Thesis Project on impact of amino acids on tumor growth
Investigate the influence of amino acids on the hyperproliferative potential of PTEN mutant cell populations in Drosophila melanogaster.
Claassen Lab Semester Project: Deep Learning for dynamic system motif detection in health and disease
Cells process external and internal biochemical signals by means of signaling cascades, which constitute complex, dynamical networks. In this project we use novel deep (machine) learning techniques to decompose such networks in their building blocks in order to elucidate the overall structure.
Christen Lab: Master Thesis Project in Experimental Systems Biology
Discovery of genetic redundancies using a targeted genome reduction approach
Hafen Group: Functional Validation of Genes Associated with Starvation Resistance
Hafen Group Student Project
Aebersold lab, Proteomics, Computational mass spectrometry and Immunology
Two bioinformatics projects available
Aebersold Lab, Proteomics, Method development in chemical cross-linking
Master thesis or semester project
Aebersold Lab, Proteomics, Method development in the field of phosphoproteomics
Master thesis or research projects available
Christen Lab: New openings for Master students
Mapping cellular core functions by transposon mutagenesis and next generation sequencing
Aebersold lab: Master thesis/semester project in Structural Proteomics
We are looking for motivated semester/master students to join our structural proteomics team at ETH to work at the interface between proteomics and structural biology.
Sauer Group, Metabolism, Systems Biology, Modelling, Bistability
Understanding history-dependence (hysteresis) of growth states in E. coli
Aebersold lab, student (master) project in phosphopeptide enrichment.
We are looking for students who would like to join project for development of the new phosphopeptide enrichent for the analysis of small sample amounts. These projects are the perfect opportunity to get hands-on experience with proteomics methods and mass spectrometry instrumentation. Prior experience with mass spectrometry is not essential, although candidates with basic knowledge of proteomics methods are preferred.
Sauer Lab, Predicting antimicrobial interactions
A project combining wet and dry bench
Sauer Group, Oxidative Stress, Metabolomics, Kinetic Modelling
A comparative study of the immediate metabolic response to oxidative stress in E. coli, Yeast and Human cells
Sauer Group, Metabolomics, Antibiotics, More of action
Identifying the target of the antimicrobial peptide Sublancin
Sauer Group, Allosteric Regulation, Metabolic Network, Kinetic Modeling
Kinetic reconstruction of E. coli central carbon metabolism with allosteric regulation
Master Thesis Project in Proteomics, Computational mass spectrometry and Immunology
We are looking for 2 highly motivated and focussed MSc students with skills in bioinformatics for immunopeptidome analysis, who will: 1) automate a computational workflow for the analysis of immunopeptidomic SWATH MS data. 2) analyze a tissue-based map of the immunopeptidome. Contact: Dr. Etienne Caron
Master Student Project in Proteomics, Epigenetics in Aging and Cancer
Epigenetic pathways are major regulators of aging and aging-associated diseases including cancer. The student will apply a recently developed mass spectrometry-based method to profile epigenetic states during aging and cancer development. Contact: Dr.Christian Feller
Claassen Group: Causality from single-cell data

Causality inference from single-cell data. Inferring cause and effect, instead of mere correlative relationships between biological variables is a challenge of interest in many biological applications. This project deals with analyzing intervention experiments with single cell readout to infer causal signaling relationships.
Sauer Group, Metabolomics, Screening, Physiology
Investigating tradeoffs between fast growth and fast switching in microorganisms
Sauer Group, Metabolomics, Modeling, Bistability
History-dependence (hysteresis) of growth states in E. coli
Sauer Lab, Metabolomics, Functional genomics
Proteome-scale enzyme discovery in yeast using nontargeted metabolomics.
Zamboni Lab, 13C metabolic flux analysis in Mycobacteria
Apply a novel framework for 13C flux analysis to co-metabolism of mycobacteria
Christen Lab, Experimental Systems Biology
Mapping essential core functions of microbial genomes by transposon mutagenesis and next generation sequencing
Claassen group, Computational Systems Biology
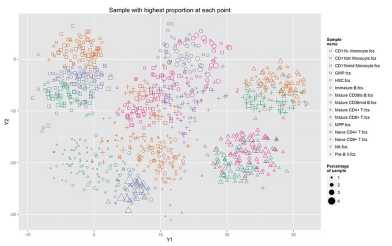
Ab initio cell type definition from high-dimensional single-cell data
Claassen group, computational systems biology
Infer structure and parameters of biochemical networks through machine learning
Sauer Group, Microbial metabolomics and modeling
Understanding the role of bacterial metabolism in the evolution of antibiotic resistance
Christen Lab, Microscopy, Asymmetric Protein Localization and Cell-Cycle Signalling in Bacteria
Investigate a novel protein localization factor
Christen Lab, Microscopy, Illuminating signaling networks in bacteria
Investigate how a secondary messenger triggers cellular heterogeneity
Christen Lab, Genomics, Quantitative Transposon Sequencing
Elucidate the core function of microbial genome by transposon insertion and NGS
Sauer Lab, Microbial metabolism, Metabolomics
A systems-level investigation of the stress-induced global metabolic adaptations in several organisms
Aebersold Lab, Proteomics, Cholesterol regulation
Analysis of cellular cholesterol regulation using targeted mass-spectrometry
