Metabolomics
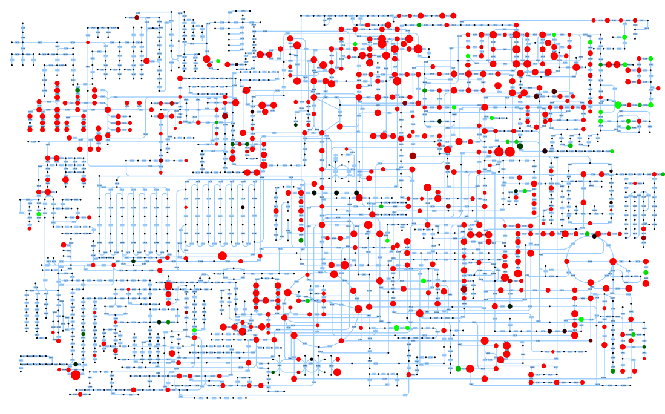
While fluxes are the functional output of metabolism, intracellular metabolite concentrations are its key constituents. In collaboration with the Zamboni lab, we develop metabolomics methods on several high-end mass spectrometers. In collaboration with Agilent Inc. we focus currently on applications of our novel high-throughput metabolomics method that quantifies 500-1000 metabolites in a given sample per minute. Our latest advance is the ability to measure intracellular metabolites automatically in real-time. With the analytical problems of data generation largely solved, the current focus is on computational data analysis and biological hypothesis generation. Several large-scale metabolomics projects, including the entire E. coli gene deletion library, are currently being finalized.
References
- Link H, Fuhrer T, Gerosa L, Zamboni N, and Sauer U (2015). Real-time metabolome profiling of the metabolic switch between starvation and growth. Nat. Methods, vol. 12, no. 11, pp. 1091–1097
- Sevin DC & Sauer U. Ubiquinone accumulation improves osmotic stress tolerance in E. coli. Nature Chem Biol. 2014 10:266-272 external pagePubmedcall_made
- Zimmermann M, Sauer U, Zamboni N. Quantification and mass isotopomer profiling of α-keto acids in central carbon metabolism. Anal Chem 2014; 86(6):3232-7 external pagePubmedcall_made
- Heux S, Fuchs T, Buhmann J, Zamboni N & Sauer U. A high-throughput metabolomics method to predict high concentration cytotoxicity of drugs from low concentration profiles. Metabolomics 8:433-443.
- Fendt SM, Büscher JM, Rudroff F, Picotti P, Zamboni N & Sauer U. Tradeoff between enzyme and metabolite efficiency maintains metabolic homeostasis upon perturbations in enzyme capacity. Mol. Sys. Biol. 2010; 13;6:356. external pagePubmedcall_made
